In December of 2024, a man in Sydney named Paul Conyngham sat across from a veterinarian who told him his dog was going to die.
Rosie was a Staffordshire Bull Terrier, Shar Pei cross he had adopted from a shelter five years earlier. Mast cell cancer. Aggressive. Surgery hadn’t stopped it. Chemotherapy hadn’t stopped it. The tumors kept returning. The vet’s estimate was months. Maybe less.
Conyngham is not a doctor. Not a veterinarian. Not a molecular biologist. He runs a machine learning consultancy in Sydney. He builds data models for a living. And when the people who were supposed to have the answers told him there were no more answers, he did something that most people would call unreasonable.
He sat down at his computer and started trying to cure her himself.
He paid three thousand dollars to have Rosie’s tumor DNA sequenced alongside her healthy cell DNA at UNSW’s Ramaciotti Centre for Genomics. He compared the two sequences and identified the somatic mutations unique to the cancer, mutations that produce abnormal proteins called neoantigens that appear only on tumor cells. He used ChatGPT to develop a treatment planning framework. He ran the mutant protein sequences through Google DeepMind’s AlphaFold to predict their three dimensional shapes. He built custom machine learning algorithms to prioritize which neoantigens would most likely trigger an immune response. The output was a half page formula describing an mRNA sequence encoding those tumor specific targets.
Then he walked that formula into a university lab and asked for help.
Pall Thordarson, director of UNSW’s RNA Institute, reviewed the analysis. He agreed to collaborate. His team synthesized the bespoke mRNA vaccine in under two months. Rachel Allavena, a professor at the University of Queensland, administered the vaccine under existing ethics approval for experimental canine treatments. Rosie received her first injection in December 2025. A booster in January 2026.
Within weeks, her tumors shrank by an estimated fifty to seventy five percent. Her energy returned. Her behavior normalized. Martin Smith, a researcher at UNSW involved in the genomic sequencing, called the result surprising and significant. Thordarson confirmed it was the first personalized cancer vaccine ever designed for a dog.
A data guy. A laptop. A dog he loved. And the refusal to accept the word no.
I watched this story on a screen in my living room, and something happened that I did not expect. I did not feel inspired. Inspiration is cheap. What I felt was recognition.
Not of the cancer. Of the posture.
I have lived with primary lymphedema since I was a child. My left leg first, swollen and heavy before I had language sophisticated enough to describe what was wrong. Then my right, years later, as if the disease wanted to make sure I understood this was not an accident but a pattern. A genetic inheritance. Autosomal dominant. Passed down through generations like a family name no one asked for.
I have been to Stanford. I have had lymphovenous anastomosis microsurgery. I have worn compression garments every day of my adult life. I have been managed. Monitored. Maintained. And at every stage, the message from the medical system has been some version of the same sentence.
This is chronic. Learn to live with it.
That sentence is a map. And I have spent decades mistaking it for the territory.
There is a map that gets handed to you early in life if you carry something the system doesn’t know how to fix. The map says: this is where the boundary is. This is what is possible and what is not. This is the edge of the known world, and beyond it there is nothing for you.
The map is drawn by good people. Competent people. Doctors who have studied the disease and understand its mechanisms. Researchers who know the literature. They are not lying to you. They are telling you what they see from inside the best map available to them.
But the map is not the territory. It never has been.
The map of NO is not a wall. It is a drawing of a wall. And there is a difference between a line on paper and a structure you cannot pass through that most people never test, because the drawing is so convincing they stop walking.
Conyngham did not stop walking.
He did not accept the map. He did not argue with the map. He did not petition the mapmakers for a better map. He walked past the edge of it and into the territory itself, carrying tools the mapmakers didn’t know how to use yet.
That is what I intend to do.
Here is what I am going to do, laid out plainly, because I have no interest in performing ambition without specifics.
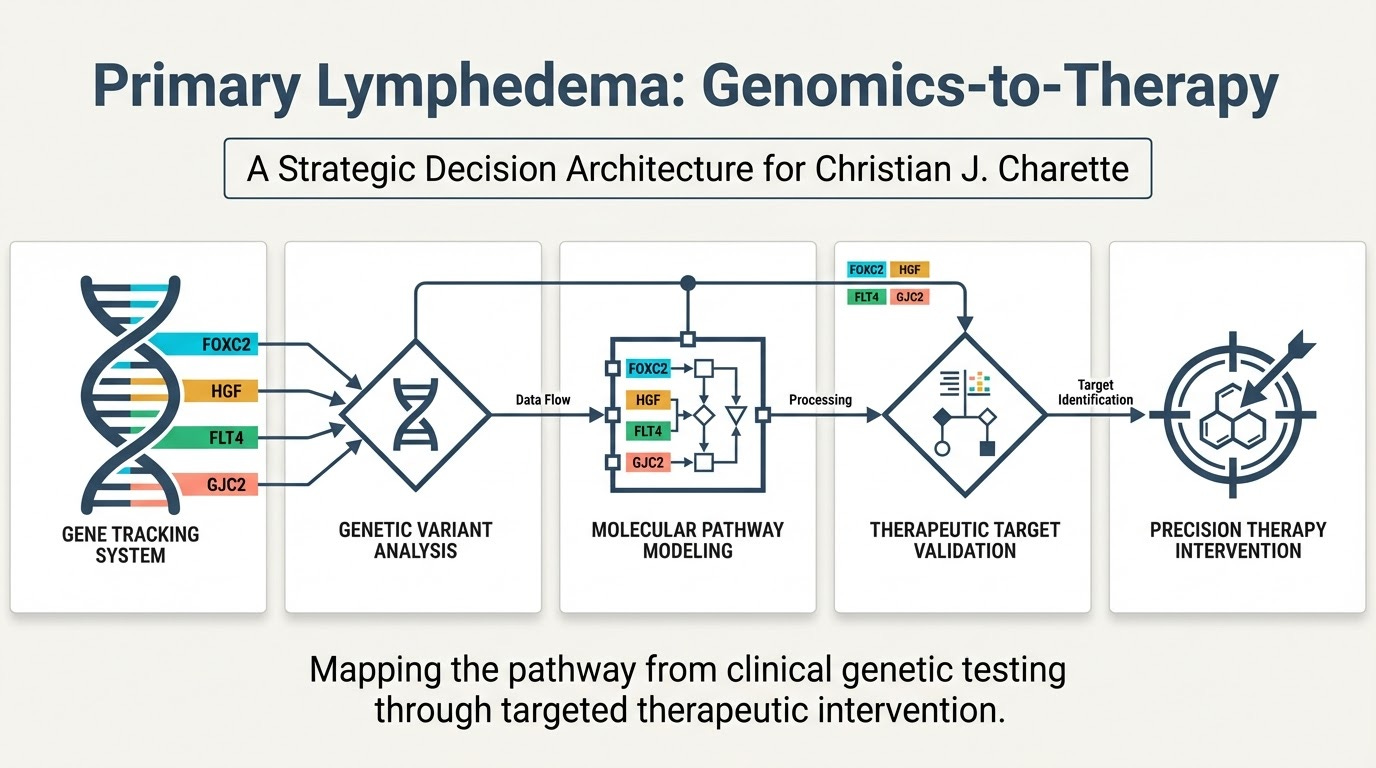
My daughter Alexis has the same disease. Same gene. Same inheritance pattern. We are two data points in the same pedigree, which means we are more valuable to researchers together than either of us is alone. Step one is ordering a clinical multigene panel for both of us. FOXC2, GJC2, HGF, FLT4. These are the four genes most commonly implicated in hereditary primary lymphedema.
The panel will return a variant report classifying each finding as pathogenic, likely pathogenic, variant of uncertain significance, likely benign, or benign. This cannot be done through 23andMe. SNP arrays do not detect the kinds of mutations that cause this disease. It requires clinical grade sequencing through a lab like Invitae, GeneDx, or Ambry Genetics. Turnaround is two to four weeks.
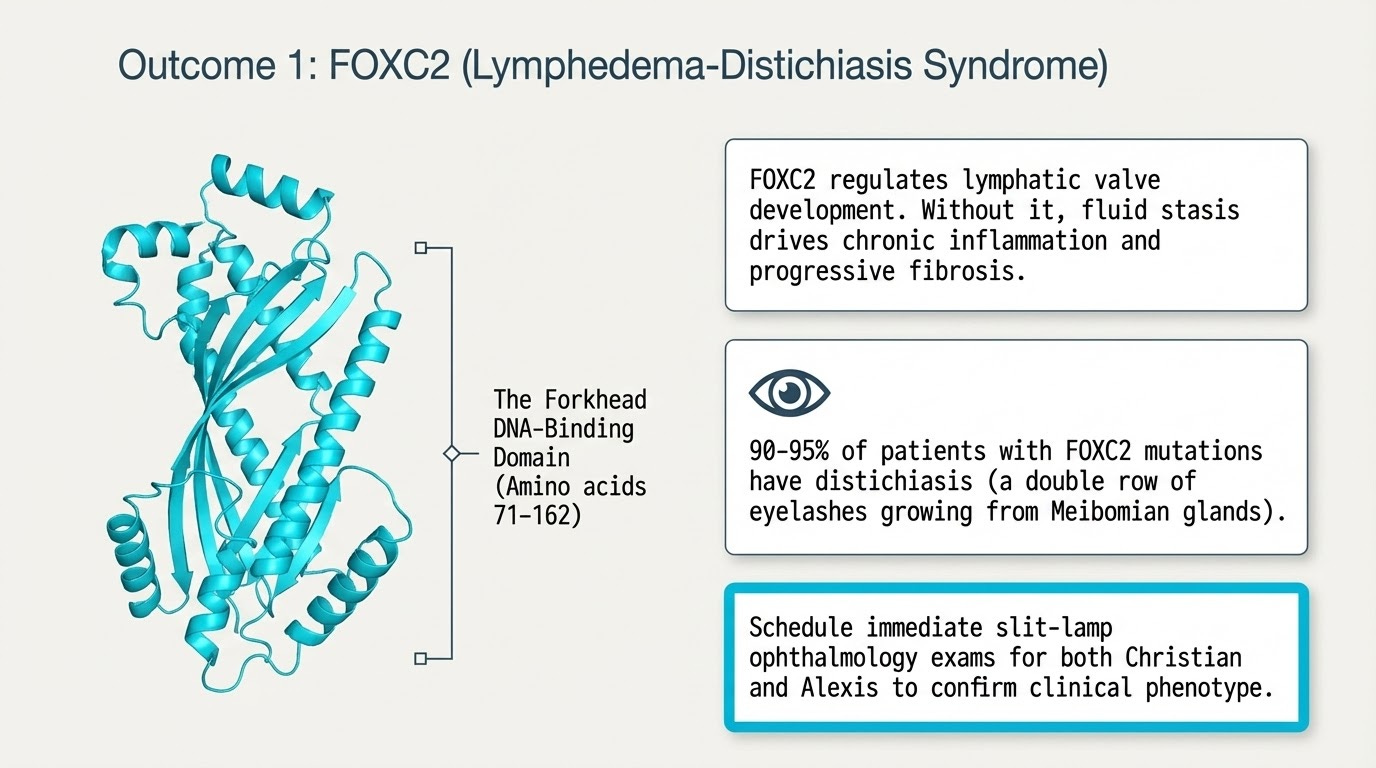
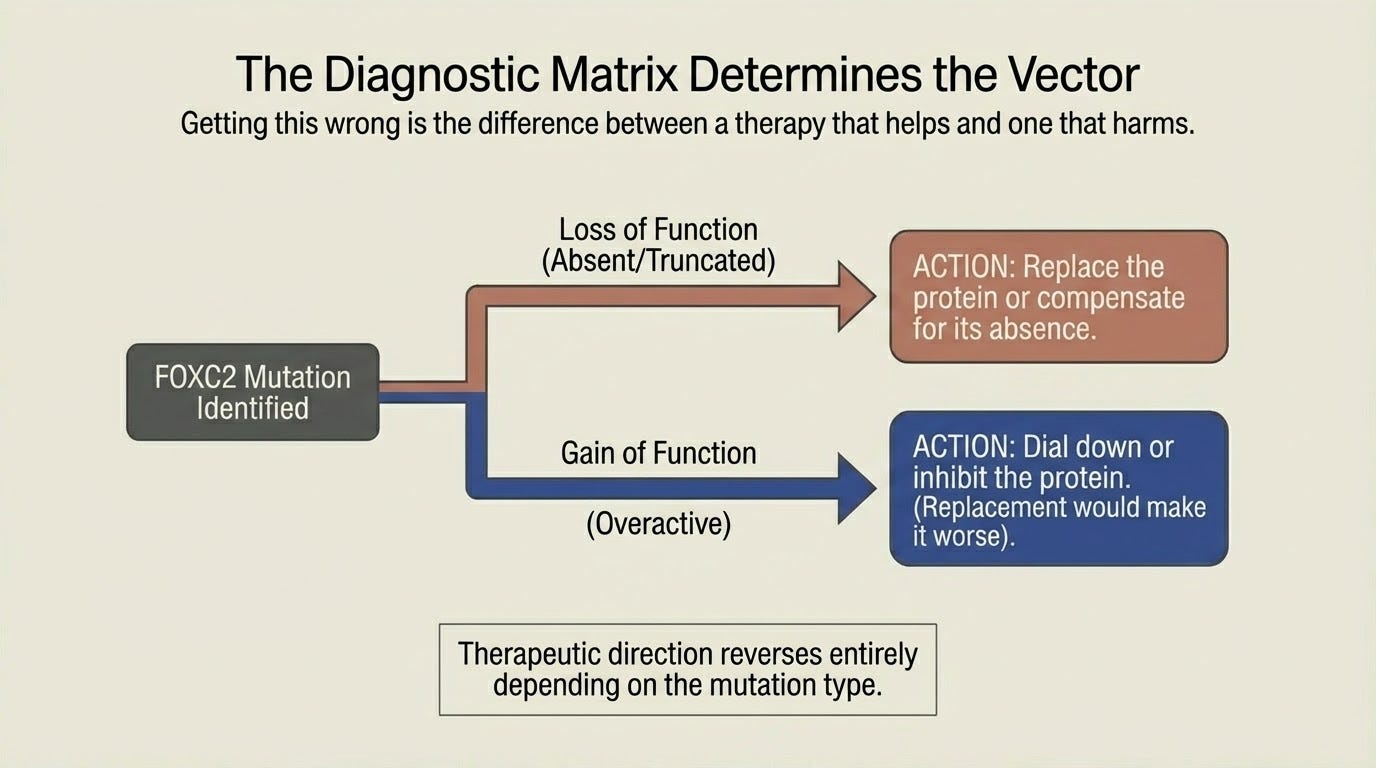
Once I have the variant, I will know which protein is broken and how it is broken. This matters more than most people realize. FOXC2, the gene most consistent with my clinical presentation, causes disease through both loss of function and gain of function mutations. The therapeutic direction reverses depending on which one you have. If the protein is absent or truncated, you need to replace it or compensate for its absence. If the protein is overactive, replacement would make things worse. You would need to dial it down. Getting this wrong is not a minor error. It is the difference between a therapy that helps and one that harms.
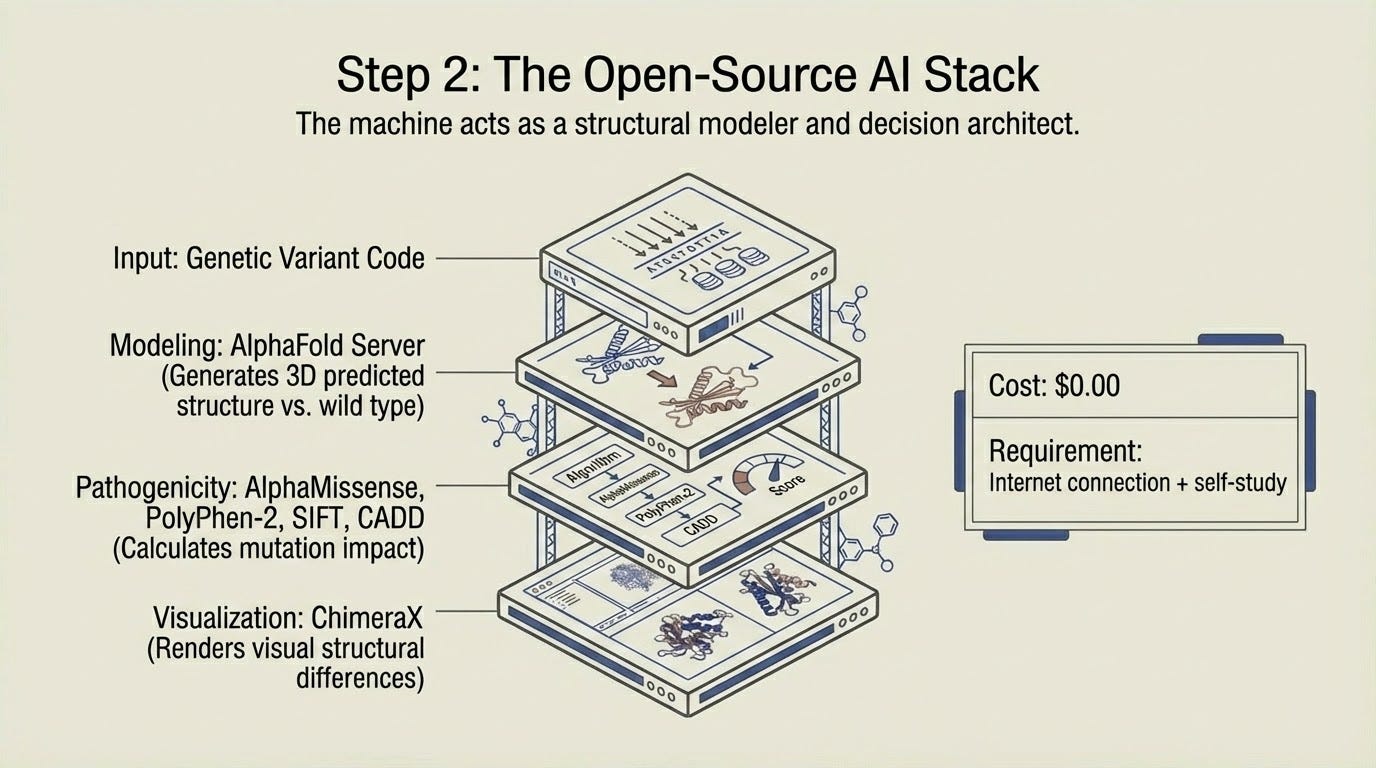
I will take the variant and submit it to AlphaFold Server to predict the three dimensional structure of my mutant protein. I will compare it against the wild type structure. I will run AlphaMissense, PolyPhen-2, SIFT, and CADD to generate pathogenicity predictions. I will visualize the structural differences in ChimeraX. All of these tools are free. All of them are available to anyone with an internet connection and the willingness to learn. The learning curve is moderate. A few weeks of self study. YouTube tutorials exist for every step.
Then I will do what Conyngham did. I will take my data and walk it into a lab.
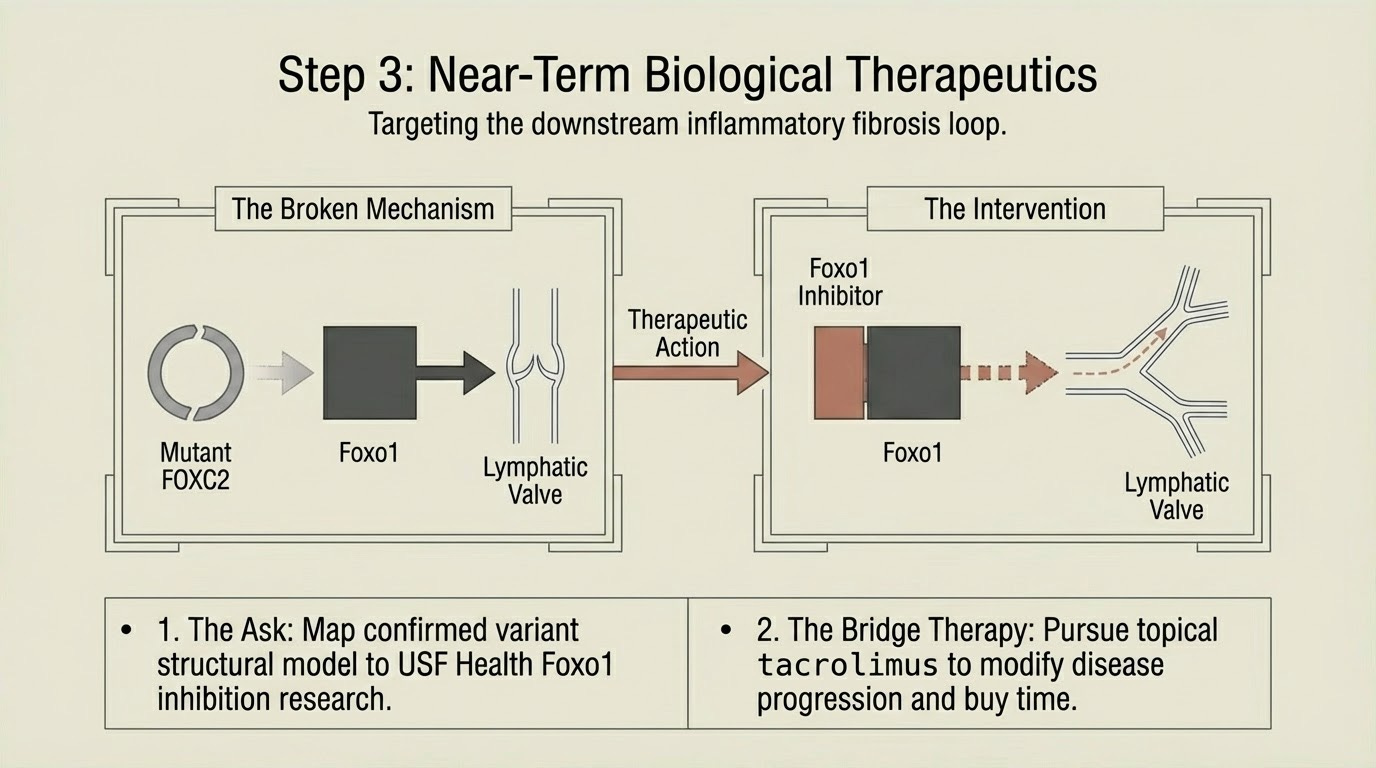
If my mutation is loss of function, the most promising near term pathway is Foxo1 inhibition. Published in the Journal of Clinical Investigation in 2021. Preclinical mouse model of human primary lymphedema. When FOXC2 cannot do its job, Foxo1 remains active and actively suppresses lymphatic valve formation. Inhibit Foxo1, remove the brake, and new valves can form even without functional FOXC2. The study demonstrated both new valve growth and repair of defective valves. There is a research group at USF Health that published this work. I intend to contact them with my confirmed variant and structural modeling and ask a specific question: does my FOXC2 variant map to your model, and would Foxo1 inhibition be predicted to compensate?
Simultaneously, I will pursue topical tacrolimus with my physician. Already identified as the most immediately actionable emerging treatment for lymphedema. It interrupts the inflammatory fibrosis loop downstream of lymphatic valve dysfunction. Not a cure. A disease modifying intervention that buys time while the genetic approaches mature.
I will assemble a research ready data package. Variant report. Structural comparison. Pathogenicity predictions. Family pedigree. Clinical history. Loss of function versus gain of function determination. This package transforms me from a patient into a collaborator. Researchers respond differently to someone who brings data than to someone who asks for help.
I will register with LE&RN, the Lymphatic Education and Research Network. I will contribute my clinical and genomic data to RARE-X, Global Genes’ shared dataset. I will submit to RDCA-DAP, the FDA supported Rare Disease Cures Accelerator platform. The point of all three is the same. I become visible to the system instead of waiting for the system to find me.
And on the frontier, the longer horizon, the tools that are coming: mRNA protein replacement therapy. The same mechanism that powered the COVID vaccines, repurposed not for immune training but for protein replacement. Deliver mRNA encoding functional FOXC2 to lymphatic endothelial cells. They produce normal protein temporarily. Valve function improves. Repeat dosing. Five to ten years from clinical availability. But the structural data I generate now is exactly what an mRNA therapy program would need as input.
CRISPR gene correction. Already demonstrated at 89% efficiency in iPSC models for FOXC2 mutations. The editing works. What doesn’t exist yet is the delivery vehicle to get it specifically to lymphatic endothelium in a living human. Seven to fifteen years out. But my confirmed variant and structural data determine which guide RNA to design. The work I do this year feeds the pipeline that could cure this disease in a decade.
Every step I just described was planned, organized, modeled, and mapped with the help of artificial intelligence. Not as a novelty. As a thinking partner. As a structural modeler. As a decision architect. As the thing that lets a therapist in private practice do work that ten years ago required an entire research department.
I need to say something about the moment we are living in, because I do not think most people have caught up to it yet.
We are in the first generation of human history where the tools of serious scientific inquiry are not locked behind institutional gates. The protein modeling software that pharmaceutical companies paid millions to access five years ago is now free and browser based. The genomic databases are open. The computational drug screening tools are open source. The machine learning frameworks that Conyngham used to prioritize his dog’s neoantigens are available to anyone who can write a line of code or, increasingly, to anyone who can describe what they want to a language model that writes the code for them.
This is not a small change. This is a phase transition.
For the entire history of medicine, the patient has been a passive recipient. You go to the doctor. The doctor tells you what is possible. You accept the map. If the map says no, you go home.
AI did not just give us faster tools. It gave us a different question. The question used to be: what will the experts do for me? Now the question is: what will I do with what is available?
That is a dignity question. And it is the most important question most people are not yet asking.
I am a therapist. I spend my professional life helping people see the maps they are living inside. Maps of who they are. Maps of what their partner is capable of. Maps of what love requires and what conflict means and what change looks like. My entire body of work is built on one insight from a Polish philosopher named Alfred Korzybski who said, in 1933, that the map is not the territory.
I have applied that insight to relationships. To attachment. To identity. To the stories couples tell about each other that calcify into certainties and slowly suffocate the people underneath them.
I never expected to apply it to my own body.
But here I am. Living inside a map that says this disease is chronic, incurable, manageable at best. A map drawn by competent people who were working with the tools available to them at the time they drew it. And I am looking at that map the same way I look at every map that crosses my desk in clinical practice.
With respect. And with the full intention of walking past it.
Paul Conyngham did not cure cancer. He cured his dog’s cancer, one specific cancer, with one bespoke intervention, built on the back of tools that did not exist three years ago. The veterinarians who told him Rosie was going to die were not wrong. They were working from the best map they had. He just refused to let their map be the last word.
I do not know if I will cure my lymphedema. I do not know if the variant will be in the gene I expect, or if the mutation type will point to the pathway I am hoping for, or if the researchers I contact will see what I see in the data. I do not know any of that.
But I know this.
The NO was never a wall. It was a map someone drew and handed me. A very good map. A carefully researched map. A map that represented the best understanding of the territory at the time it was made.
But maps age. Tools change. Territories shift. And sometimes a person sitting in their living room, watching a story about a man and his dog, realizes that the edge of the known world is not where they were told it was.
It is wherever you decide to stop walking.
I am not stopping.